
Analyzing Pathways from PTMs: A Guide
Lucia Williams, Nagashree Avabhrath, Mikhail Ukrainetz, Madison Moffett, Grant Smith, Mark Grimes
RawDataProcessing.RmdPurpose
This vignette intends to help users produce a matrix with PTM names as row names (e.g. FYN p Y411) and numeric data in all the columns, which can be used as input to the PTMsToPathways functions.
Mass spectrometry data output will vary depending on the experimental
design, source of data, and software used to process the raw spectra. R
supports many file types and can automatically convert them into a data
frame. For example, read.csv() will take a csv file and
convert it into a data frame (read.csv() is a variation of
read.table()). We start with data in tab-delimited
spreadsheet format.
Naming conventions
In this vignette, we use the following shorthand conventions when describing PTMs based on the modifications present in the example data set. Your data will dictate the names of modifications.
- Gene.Name = The HUGO Gene Name is used to identify the protein/gene
- Phosphorylation = “p”
- Lysine acetylation = “ack”
- Lysine methylation = “kme”
- Arginine methylation = “rme”
- Ubiquitination = “ubi”
Data input
Investigators will need to identify which columns contain key information for analysis of PTMs. The example raw data file for this vignette (downloadable here, or load directly into R using the commands below) contains only phosphorylation sites.
The key columns are:
-
Amino Acid: the modified amino acid, such as S or T -
Positions Within Proteins: the amino acid number in the protein sequence - Ambiguous modification sites: multiple possible positions separated
by
; -
Modification Type: the PTM class, such as phosphorylation
R converts spaces in column names to periods, so the relevant columns after import are:
-
genes="AllGeneSymbols" -
positions="Positions.Within.Proteins" -
aa="Amino.Acid" -
modification="Modification.Type"
For input to PTMsToPathways, it important that the row names are PTM
identifiers and that the columns represent experimental conditions. The
numeric values are the mass spectrometer output, and NAs represent
missing data rather than zeroes. Ambiguous PTMs, where a PTM could match
several proteins, are separated by semicolons (for example,
"AARS ubi k747; AMBLIL p U123").
To prepare data for input to PTMsTo pathways, we first read in the
data file. To use your own data file, replace file_path
variable with your own path to file, as in the commented line below.
# file_path <- "path/to/your/file.txt"
file_path <- system.file("extdata", "phospho_cleaned_mapped.txt",
package = "PTMsToPathways")
newphos <- utils::read.table(file_path, sep = "\t", skip = 0, header = TRUE,
blank.lines.skip = T, fill = T, quote = "\"", dec = ".",
comment.char = "", stringsAsFactors = F)
dim(newphos)
>> [1] 933 170First remove internal control rows (reverse sequences).
Many investigators inspect data in Microsoft Excel, which can export
tab- or comma-delimited files. Unfortunately, Excel can silently convert
some gene names into dates when they appear in a cell by themselves. We
reverse that with the helper function fix.excel(). If there
are dates in the AllGeneSymbols column, use:
newphos$AllGeneSymbols <- sapply(newphos$AllGeneSymbols, fix.excel)Next, identify the columns that will be used to name the PTM
headercols <- c("AllGeneSymbols", "Amino.Acid",
"Positions.Within.Proteins", "Modification.Type")Make a separate data frame using these columns.
newphos.head <- newphos[,headercols]We provide another helper function, name.peptide(), to
handle ambiguous modification sites (a modification site whose peptide
sequence is the same in more than one protein) separated by “;” or
another separator.
newphos.head$Peptide.Name <- mapply(
name.peptide, genes = newphos.head$AllGeneSymbols,
sites = newphos.head$Positions.Within.Proteins, aa = newphos.head$Amino.Acid)Replacing Patterns
If the list of PTMs possesses symbols that are unnecessary such as “AARS ~K ubi K747”, this command will remove all strings included in the “patterns” vector from the rownames. Any pattern can be chosen so long as the user ensures that any special character (such as $ or @) have a “\” in front of them.
Data columns
Next we identify the data columns, which contain the string “Intensity.” The example data file is from a multi-PTM study and the data in this table are from just the phosphorylation pulldown (other tables are for other PTM types). The optimal pulldown columns are straightforward to identify by the pulldown strings present in the sample names, “pTyr” in this case. They are also identifiable by zooming out and looking at the patterns of missing data, the optimal pulldowns, as a group, have the least missing data. The second step is to select colums that have “pTyr.”
These data have technical replicates, which means that the same samples were run twice. Due to the stochastic selection of peptides for detection, the pattern of missing values is slightly different between technical replicates. We therefore merge the technical replicates taking the value of either replicate where it’s missing in the other, and averaging values detected in both, using the PTMsToPathways function merge2cols().
Key for column names
C = Crizotinib
D = DMSO
E = Erlotinib
Pr = PR171
For Intensity columns
C1-1: Crizotinib biological replicate 1- technical replicate 1
C1-2: Crizotinib biological replicate 1- technical replicate 2
C2-1: Crizotinib biological replicate 2- technical replicate 1
C2-2: Crizotinib biological replicate 2- technical replicate 2
C3-1: Crizotinib biological replicate 3- technical replicate 1
C3-2: Crizotinib biological replicate 3- technical replicate 2
… & similar for other drugs
phosdata <- newphos[,grep("Intensity", names(newphos))]
phosdata <- phosdata[,grep("pTyr", names(phosdata))]Now simplify column names:
Make zero into NA, which it is. (Note that this may not apply if you are confident that zero means actual zero, which is possible with certain technical advances like DIA.)
Define technical replicates:
tr1.opt <- names(phosdata)[grep(".1", names(phosdata), fixed=TRUE)]
tr2.opt <- names(phosdata)[grep(".2", names(phosdata), fixed=TRUE)]Use merge2cols() to average technical replicates. This
function ignores NA values in either column and takes the average in the
case where there are two values.
phosdata.merged <- data.frame(matrix(nrow=nrow(phosdata), ncol=18))
for(i in 1:length(tr1.opt)) {
phosdata.merged[,i] <- mapply(merge2cols,
colv1 = as.numeric(phosdata[, tr1.opt[i]]),
colv2=as.numeric(phosdata[,tr2.opt[i]]))}
names(phosdata.merged) <- sapply(tr1.opt, function(x){
substr(x, start=1, stop=nchar(x)-2)
})Merge with header:
phosdatafile <- cbind(newphos.head, phosdata.merged)This file could be saved for reference using
write.table():
write.table(phosdatafile, file = "phosdatafile.txt",
row.names = FALSE, sep = "\t")For subsequent steps, we make a dataframe withjust the data with individual PTMs as rownames:
rownames(phosdatafile) <- phosdatafile$Peptide.Name
phosdata.df <- phosdatafile[,6:23]Log base 2 transformation improves clustering.
log2phosdata <- log2(phosdata.df)The Creating Networks
vignette show how to use the functions provided in PTMsToPathways to
analyze data stored in a variable called ptmtable.
ptmtable <- log2phosdataOptional data processing steps
For experiments where treatment with drugs is compared to control samples, adding treatment/control ratios as additional data column can improve clustering. This optional step adds dimensions to the data set that enhance focus on the changes in response to drug treatments.
Simplify column names first:
names(phosdata.df) <- sapply(names(phosdata.df), function(x){
paste(unlist(strsplit(x, "SEPTM_pTyr"))[1],
unlist(strsplit(x, "SEPTM_pTyr"))[2], sep = "")
})Explore using ratios where control=rowMeans (D1, D2, D3):
H3122control <- rowMeans(phosdata.df[, names(phosdata.df)
[grep("H3122.D", names(phosdata.df))]],
na.rm=TRUE)Change NaN to NA
H3122control[is.nan(H3122control)] <- NA
PC9control <- rowMeans(phosdata.df[, names(phosdata.df)
[grep("PC9.D", names(phosdata.df))]],
na.rm=TRUE)
PC9control[is.nan(PC9control)] <- NACalculate treatment/control ratios
# H3122 cells
H3122.C1.ratio <- phosdata.df$H3122.C1/H3122control
H3122.C2.ratio <- phosdata.df$H3122.C2/H3122control
H3122.C3.ratio <- phosdata.df$H3122.C3/H3122control
H3122.PR1.ratio <- phosdata.df$H3122.PR1/H3122control
H3122.PR2.ratio <- phosdata.df$H3122.PR2/H3122control
H3122.PR3.ratio <- phosdata.df$H3122.PR3/H3122control
# PC9 cells
PC9.E1.ratio <- phosdata.df$PC9.E1/PC9control
PC9.E2.ratio <- phosdata.df$PC9.E2/PC9control
PC9.E3.ratio <- phosdata.df$PC9.E3/PC9control
PC9.PR1.ratio <- phosdata.df$PC9.PR1/PC9control
PC9.PR2.ratio <- phosdata.df$PC9.PR2/PC9control
PC9.PR3.ratio <- phosdata.df$PC9.PR3/PC9controlPut these columns in a data frame:
phos_ratios <- data.frame(H3122.C1.ratio, H3122.C2.ratio, H3122.C3.ratio,
H3122.PR1.ratio, H3122.PR2.ratio, H3122.PR3.ratio,
PC9.E1.ratio, PC9.E2.ratio, PC9.E3.ratio,
PC9.PR1.ratio, PC9.PR2.ratio, PC9.PR3.ratio)Check (should be TRUE):
Make limits to unweight extreme values. This has been shown to improve clustering, and a ratio of 1000 is biologically not really functional different than a ratio of 100.
hi.ratio <- which(phos_ratios >= 100, arr.ind = TRUE)
low.ratio <- which(phos_ratios <= 1/100, arr.ind = TRUE)
phos_ratios.lim <- replace (phos_ratios, hi.ratio, 100)
phos_ratios.lim <- replace (phos_ratios.lim, low.ratio, 1/100) log2 transformation improves clustering:
phos_ratios.lim.log2 <- log2(phos_ratios.lim)
phosdata_plus_ratios <- cbind(log2phosdata, phos_ratios.lim.log2)And plot to check:
boxplot(phosdata_plus_ratios)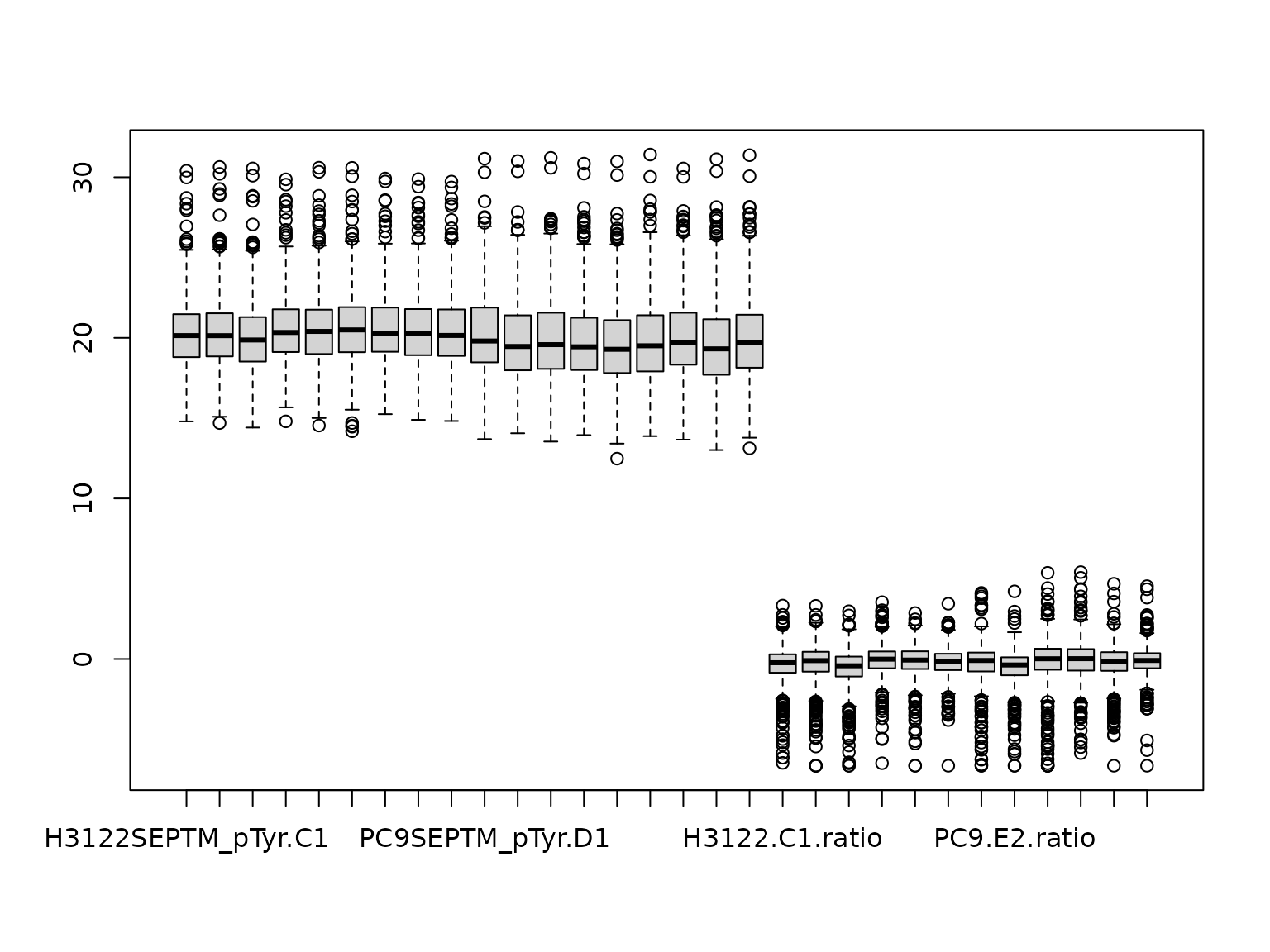 And do one more check:
And do one more check:
This can be used as the example ptmtable for subsequent testing.
# ptmtable <- phosdata_plus_ratiosCombining data from multiple PTM experiments
For experiments involving multiple PTMs, or if investigators wish to combine several data sets, the data can be combined. For example, pulldowns were made to isolate acetylated and ubiquitinated peptides from the same experimental samples. Combining the data is simply repeating the above steps using the correct optimum data columns, then making the column names the same, and binding all the rows together.
Suppose you had acetylation data in ackdata.df and
ubiquitination data in ubidata.df, both formatted as above
for phosphorylation data in phosdata.df. You could combine
them as follows.
First, make sure the column names are the same:
kgp <- phosdata.df
kga <- ackdata.df
kgu <- ubidata.df
names(kgp) <- sapply(names(kgp), function(x){
paste(unlist(strsplit(x, "_pTyr"))[1], unlist(strsplit(x, "_pTyr"))[2],
sep = "")
})
names(kga) <- sapply(names(kga), function(x){
paste(unlist(strsplit(x, "_AcK"))[1], unlist(strsplit(x, "_AcK"))[2],
sep = "")
})
names(kgu) <- sapply(names(kgu), function(x){
paste(unlist(strsplit(x, "_Ubi"))[1], unlist(strsplit(x, "_Ubi"))[2],
sep = "")
})
identical(names(kgp), names(kga)) # Check TRUEThen rbind them:
ptmdata <- rbind (kgp, kga, kgu) Reorder:
This optional step improves clustering in our hands:
log2ptmdata <- log2(ptmdata)Finally, this dataframe could be used as the example
ptmtable for the P2P functions.
# ptmtable <- log2ptmdata